 Gnotobiology isn't new, but the dramatic expansion of microbiome research has outpaced the development of a common nomenclature within the gnotobiotic research community. Many different, and sometimes confusing, terms are used when referring to different gnotobiotic concepts. To help, here's an overview of the most commonly used gnotobiotic rodent model concepts with examples of how these concepts are applied in basic research and drug discovery.
Gnotobiology isn't new, but the dramatic expansion of microbiome research has outpaced the development of a common nomenclature within the gnotobiotic research community. Many different, and sometimes confusing, terms are used when referring to different gnotobiotic concepts. To help, here's an overview of the most commonly used gnotobiotic rodent model concepts with examples of how these concepts are applied in basic research and drug discovery. Gnotobiology means the study of "known life." Hence, a "gnotobiotic animal" is a host animal harboring only defined (known) microbial life, if any. Gnotobiotic animal models may be either germ-free or be known to carry only a defined set of microorganisms. Somewhat confusingly, "gnotobiotic" is sometimes used in a broader sense to cover mice or rats that were initially germ-free and subsequently colonized, or associated, with a complex microbial community. These communities may be characterized to some extent, but are rarely fully defined, as this would require in-depth analysis of all bacteria, viruses, fungi, protozoa and more.
Conceptual Overview of Gnotobiotic Rodent Models
Concept | Also Known As | Definition | Why Use This Concept |
| Reductionist approaches | |||
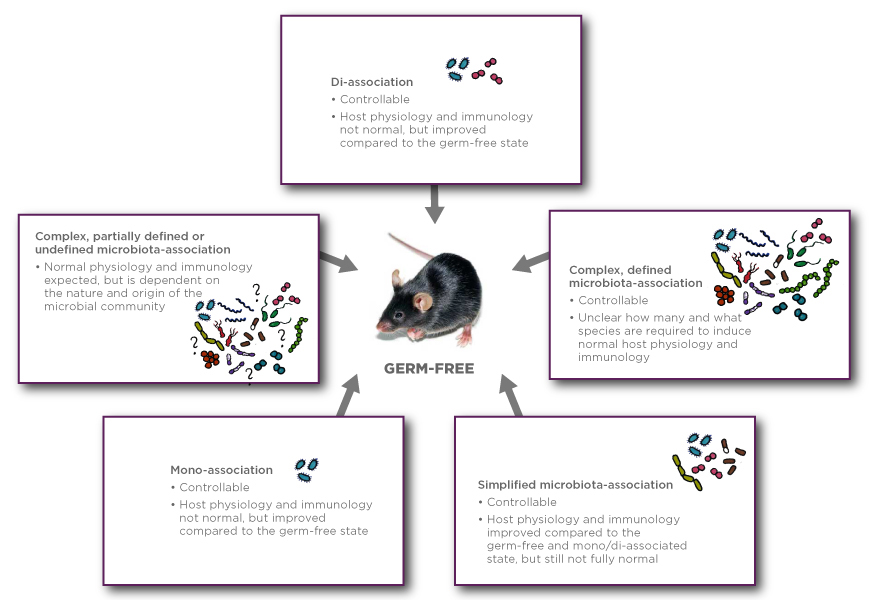
| Germ-free | Axenic, sterile | Germ-free rodents lacking all known organisms (bacteria, fungi, viruses, parasites) except for endogenous retroviruses. The germ-free state is determined within the limitations of the currently available detection methods. | Germ-free rodents are considered the gold standard as reliable and reproducible tools for microbiota-association studies and as founders for gnotobiotic or standardized colonies or study cohorts. Germ-free rodents are additionally crucial as controls in association studies (also called fecal microbiota transplantation). |
| Mono-associated | Monoxenic, mono-colonized | Rodents harboring just one organism, e.g. Escherichia coli, Segmented Filamentous Bacteria, or Bacteroides thetaiotaomicron | To study the function and effects of a particular organism, e.g. induction of the immune system by the organism1 |
| Di-associated | Dixenic, di-colonized, dual-associated, bi-associated, bi-colonized | Rodents harboring just two organisms, e.g. B. thetaiotaomicron in conjunction with Faecalibacterium prausnitzii. The concept may be expanded to tri-associated (trixenic), tetra-associated and so on. | To study synergy, interaction, and adaptation between two organisms and the host2. To facilitate the colonization with extremely oxygen sensitive bacteria by initial inoculation with an aerotolerant species that prepare an anaerobic enviroment3. |
| Simplified microbiota-associated | Defined flora/microbiota-associated/colonized | Rodents harboring a limited number of known (defined) organisms, e.g. the mouse-derived Altered Schaedler Flora or communities with a limited number of human-derived bacteria or from other target species. Such limited consortia typically consist of 5-15 species and may be procured from microbial repositories. | For a tractable, time stable, and reproducible model system to study, for example: |
| Holistic approaches* | |||
| Mouse/rat microbiota-associated with: - healthy microbiota - dysbiotic microbiota - disease model-prone microbiota - wild-sourced microbiota | Complex microbiota-associated, SPF-colonized, conventionalized | Rodents harboring another rodent microbiota than their endogenous microbiota. Typically generated by inoculating germ-free mice or rats, but other approaches such as cross-fostering and embryo transfer may also be used. | Phenotype transfer for proof-of-concept hypothesis testing before moving to human patient samples that may more difficult to access8. To optimize disease model readout and pursue the 3Rs by creating high-responder cohorts9. To improve translation to humans by creating models with a naturalistic wild mouse microbiota and hence, an improved immune response10. For establishing rodents with a standardized microbiota to overcome reproducibility issues due to variation in the microbiome over time or between locations. |
| Human microbiota-associated with: - healthy microbiota - dysbiotic microbiota - patient microbiota | Humanized, complex microbiota-associated, human flora-colonized, microbial xenoengraftment | Rodents colonized with human intestinal microbiota, most often from feces. The proportion of the donor microbiota that is successfully established in the rodents varies and is often smaller than it is for mouse/rat microbiota-associated mice/rats. The proportion that establish in the animals is stable over time and generations under protected conditions11. Human microbiota-associated mice should ideally be validated for study of specific immune parameters since immunomodulation of the host is altered in these mice. | For a translational and predictive model to study for example: |
* The holistic approaches are strictly speaking not gnotobiotic unless all members of the complex community are known and precisely defined. While this is rarely the case, these models are nevertheless often referred to as gnotobiotic since they require gnotobiotic or clean IVC housing technology.
















.jpg)

.jpg)
.jpg)
.jpg)
.jpg)





.jpg)


.jpg)
.jpg)




.jpg)





.jpg)

.jpg)





