While recent advances in gene editing accelerated the migration of models to a pure C57BL/6 background, researchers should remain aware of potentially confounding results from genetic variation between C57BL/6 substrains.
Problems with Early ES Cell Lines
The vast majority of contemporary mouse studies use the C57BL/6 inbred strain, which is an excellent model for applications such as metabolism, behavior, immunology, and oncology. However, early knockout models were produced on a different genetic background, using embryonic stem (ES) cells derived from substrains of inbred 129 mice.These 129 strains are quite distinct from C57BL/6 mice, are not suitable for many studies, and suffer from poor breeding performance.
Extensive backcrossing is commonly employed to transfer knockout lines to a congenic C57BL/6 background, but this process is both time consuming and expensive. Furthermore, backcrossing cannot completely remove tightly linked 129-derived alleles flanking the targeted locus. This may result in a region of around 400 contaminating 129 donor genes, with potentially significant impacts on mouse phenotype and interpretation of results1.
Emergence of the C57BL/6 ES Cell
An obvious way to address these issues is to create mouse GEMs on a pure C57BL/6 background. For many years this was problematic; C57BL/6-derived ES cells were considered intractable, being both unstable and inefficient compared to 129-derived ES cells. However in the past decade, efficient C57BL/6 ES cells have been created from C57BL/6NTac mice which circumvent these limitations (Figure 1).These C57BL/6NTac ES cells are reported to show "better growth properties and morphology compared to C57BL/6J ES cells2." Several of these, including the JM8, C2, and VGB6 ES cell lines2,3, and Taconic's proprietary, highly germline competent ES cells, have become powerful and versatile tools for creating knockouts and knock-ins on a pure C57BL/6 background.
ES Cell Line | Background |
| JM8 | C57BL/6NTac |
| JM8.F6 | C57BL/6NTac |
| JM8.N4 | C57BL/6NTac |
| JM8A3 | C57BL/6NTac |
| JM8A3.N1 | C57BL/6NTac |
| JM8A1.N3 | C57BL/6NTac |
| C2 | C57BL/6NTac |
| D4 | C57BL/6NTac |
| WB6d | C57BL/6NTac |
| VGB6 | C57BL/6NTac |
| iTL IC1 | C57BL/6NTac |
Figure 1: Strain backgrounds of commonly-used C57BL/6 ES cells
Predominance of C57BL/6NTac ES Cells in Modern GEMs
C57BL/6NTac-derived ES cells have become the most common tool for the generation of knockouts and knock-ins on a pure C57BL/6 background. These are the cells of choice for most large scale knockout generation and phenotyping projects worldwide, including:As of August 2016, the IMPC reported 22,277 mouse lines derived from C57BL/6NTac ES cells.
By generating large numbers of knockout lines on a single defined genetic background, these projects aim to uncover the function of every single coding gene in the mouse genome using a high-throughput approach. The IMPC recently identified 410 genes out of 1751 knockout lines which are lethal during embryonic development5. Such studies are commendable and demonstrate the power of C57BL/6NTac-derived ES cells.
Not All C57BL/6 Mice are the Same
The C57BL/6NTac strain is but one of a large family of related, but genetically distinct, inbred mice which are often referred to collectively as "C57BL/6". Different substrains of C57BL/6 mice possess unique genetic characteristics that should be considered in study design and in the generation and breeding of genetically engineered animal models.The two main branches of this family tree, C57BL/6J and C57BL/6N, diverged in 1951; there are numerous genetic variants between these two strains which could subtly affect experimental outcomes.
For example:
- Most C57BL/6J mice carry a mutation in the nicotinamide nucleotide transhydrogenase gene (Nnt), which impairs insulin secretion and causes redox abnormalities.
- All C57BL/6J mice have a mutation in the Nod-like receptor pyrin domain containing 12 (Nlrp12) gene, which impairs innate immune responses and cytokine production.
- C57BL/6N mice experience a mild form of retinal degeneration due to the Crb1rd8 mutation, and have impaired responses to amphetamines due to a mutation in Cyfip2.
The C57BL/6N branch, from which C57BL/6NTac mice are derived, are more closely related to each other than they are to C57BL/6J mice. This does not imply that they are the same, however.
While early reports concluded that all C57BL/6N substrains were genetically identical (based on limited single nucleotide polymorphism (SNP) panel testing), more recent studies have identified considerable genetic variation within C57BL/6 substrains (Figure 2)6. This known variation at the SNP level likely underrepresents the true genetic variability amongst C57BL/6 mice. Genome sequencing of the various substrains is required to characterize the full extent of functional genetic variation in C57BL/6 substrains.
| C57BL/6NTac | C57BL/6NCrl | C57BL/6J | |
| C57BL/6NTac | 0 | 287 | 464 |
| C57BL/6NCrl | 287 | 0 | 345 |
| C57BL/6J | 464 | 345 | 0 |
Figure 2: Genetic diversity of C57BL/6 mice. Shown are the number of SNPs with variation between indicated C57BL/6 substrains6. Data accessed from the Mouse Phenome Database.
Genetic variation between substrains can substantially impact studies using knockouts on the C57BL/6N background. For example, a recent study identified immune deficiencies associated with a Dock2 loss of function mutation specific to C57BL/6NHsd mice; this incidental mutation had led to erroneous conclusions when studying a Siae knockout line on that background7. We do not yet fully understand the level of genetic diversity within C57BL/6N mice, but there are likely more instances where non-synonymous genetic variants or altered regulatory elements exist between C57BL/6N substrains.
Using Mice Derived from C57BL/6NTac ES Cells
If you are using a genetically engineered mouse created from a C57BL/6NTac ES cell, should you be concerned about variation between C57BL/6 substrains?The point of creating a knockout in a "pure C57BL/6" background is to avoid issues of genetic variation and the expense of backcrossing. Not all C57BL/6 mice are identical, and ES cells derived from C57BL/6NTac mice will have some level of genetic variability compared to other C57BL/6 mice, including other C57BL/6N substrains.
The exact variability between these strains and the associated functional consequences are unknown, so there is no way to predict how breeding between substrains could affect your studies.
There is a real possibility that segregating allelic differences within experimental cohorts derived from mixed C57BL/6 strains could complicate both the interpretation and reproducibility of subsequent studies, as well. The example of the Dock2 mutation in one C57BL/6N substrain demonstrates how such variability could hinder the success of a knockout project7.
Maintaining Modified C57BL/6 Lines
Always maintain C57BL/6 GEM lines on a single, characterized inbred substrain background.According to geneticist Dr. Sara Wells of MRC Harwell, C57BL/6 substrains should be considered genetically distinct. MRC Harwell, a member of the IMPC, maintains their mouse lines on a pure C57BL/6NTac background.
To accomplish this, they breed all mouse lines derived from JM8 ES cells to genetically-matched C57BL/6NTac mice and use C57BL/6NTac mice as wild type controls. Recognizing the importance of this issue for the research community, MRC Harwell provides recommendations and materials to support proper genetic quality when distributing GEM lines from their repository.
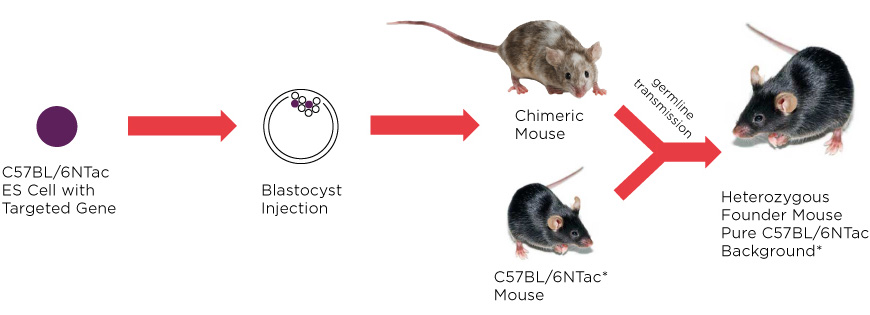
Figure 3: C57BL/6NTac ES cells provide the platform to derive GEM lines on a pure C57BL/6 background. The genetic background of ES cell clones provides 50% of the genetic makeup of the founder mice in resulting knockout lines. Mating chimeras with C57BL/6NTac mice will result in heterozygous founder mice on a defined C57BL/6NTac genetic background. Subsequent mating to C57BL/6NTac mice will ensure that allelic variants related to other C57BL/6 substrains are not introduced into your colony.
Best Practices
When planning to use a C57BL/6 knockout mouse in your study, adhere to these five best practice guidelines to avoid confounding results:- Know which background strain and substrain of ES cells were used to generate your model.
- Understand that C57BL/6 mice are not equivalent and that variation exists between C57BL/6J and C57BL/6N, as well as within C57BL/6N substrains.
- Avoid introducing genetic variation into your knockout mouse line through breeding and maintenance of the line.
- Control for genetic variation that may have been introduced into your knockout line and verify genetic background by SNP testing.
- Only use wild type controls that are as genetically identical as possible.
 Download the Taconic Biosciences' White Papers:
Download the Taconic Biosciences' White Papers:- Promise and Pitfalls: An Updated View of CRISPR/Cas9 for Genetic Engineering in Mice and Rats
- Disaster Preparedness: The Choices and Elements to Consider When Selecting Lines to Cryopreserve
- The Advantage and Application of Genetically Engineered Mouse Models for Predicting Human Efficacy and Metabolism
















.jpg)

.jpg)
.jpg)
.jpg)
.jpg)





.jpg)


.jpg)
.jpg)




.jpg)





.jpg)

.jpg)




